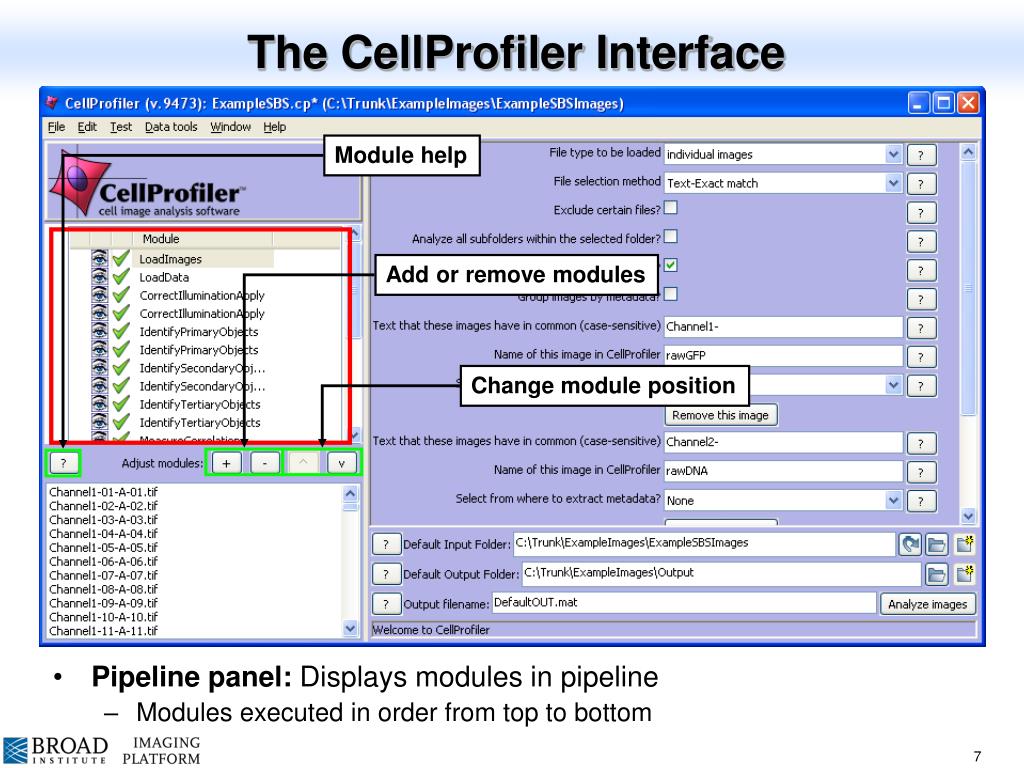
These examples show the simplest and most common cases for loading data into CellProfiler - cases where you are working with 2D images and have either one channel or a full set of matching channels for each field of view.Įxample 1: Load and configure an image set consisting of three channels saved as separate images. The first step to to start working with you images is to drag and drop your images/image folders onto the “Drop files and folders here” pane in the Images module, from there it depends on the type of images you are working with, in the next section we provide numerous examples of types of data you might be working with, while the list is extensive it is not exhaustive. This approach makes metadata extraction and downstream data analysis easier, as well as easier to find the exact file you need later. You may want to include relevant information within the filenames separated by spacers such as a dash or underscore (but NOT a space) - an example could be CellTypeA-TreatmentX_Day5-channel1.tiff. Keep in mind that the name of the files and folders can be used as metadata.Avoid special characters, such as periods, parenthesis, or slashes.Instead, separate words using either capital letters (MyData), dashes (My-Data), or underscores (My_Data). Avoid using spaces in filenames and folder names.An underappreciated yet important aspect of any image analysis project is choosing what to name your images and the folders that contain them. We encourage you to consult the Help manual the Images module section provides more details regarding digital images and file formats. Preparing your data for import into CellProfilerĬellProfiler can work with many file formats. It is also often used when running CellProfiler on a cluster or in the cloud in order to break one large experiment into many easier to analyze pieces (see our blog post on Distributed CellProfiler). This approach is helpful for many different experimental setups including Z-projecting images and analyzing time-lapse data. The Groups module splits the images into collections or subsets that can be analyzed independently. different channels from a field of view). This module is also where you can specify the type of image that is being loaded, and configure image sets (e.g. The naming can be assigned based on the filename, file extension, type of image, file directory, or image metadata. The NamesAndTypes module is where you will create names for the images to be used during the later analysis modules. The information can be used in later modules to identify the image, to create categories to group a set of images or to obtain image information (such as genotype or compound treatment) that can be stored with the results of your image analysis. The Metadata module allows you to extract information about the files. The Images module is where you will add the individual files or folders containing the images you wish to analyze this module also allows you to optionally filter the set of files to use during the analysis. time-lapse movie, z-stack or plate) Introduction An overview of the input modules (0:00) DAPI and GFP images), and an “image group” is a collection of image sets that should be processed independently from each other (e.g. In CellProfiler, an “image set” is the collection of channels representing a single field of view (e.g. Some examples required a modification of the source images, here you can access all the files used in the tutorial. For each example the source of the image sets is provided. The timestamps in this document correspond to the video tutorial. This written tutorial is accompanied by a video tutorial that can be found here. In this written tutorial, we’ll go over some best practices and examples of loading many different data types into CellProfiler. Z-stacks, time series, image masks), but it’s important to input them carefully so that CellProfiler can parse them correctly. CellProfiler can load many different data types (e.g. These modules are crucial for any CellProfiler pipeline because they define how images are loaded and organized in CellProfiler for downstream analysis.  
Most of us, however, have images with multiple channels or more complex image metadata that you need to tell CellProfiler how to identify.The tutorial aims to introduce you to four modules in CellProfiler Images, Metadata, NamesAndTypes, and Groups (collectively known as the Input modules). If you are working with single-channel images, you can just drag a few images into CellProfiler and start making your pipeline.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
